Researchers are amassing data to design precision medical treatments and prevent the associated health risks of the common condition.
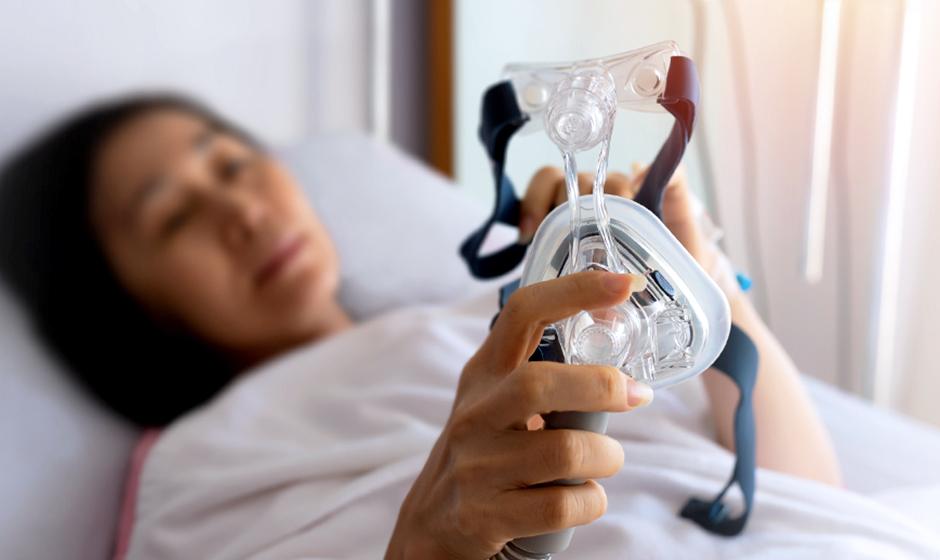
Sleep apnea can be a pain in the neck for sufferers and their significant others. It can also increase the risk of developing other harmful health conditions. A new National Institutes of Health-funded study is delving deeper into individuals’ biological traits using state-of-the-art genetic and epigenetic sequencing, along with other approaches, in an effort to improve and extend the lives of people with obstructive sleep apnea (OSA).
Led by Vancouver Coastal Health Research Institute researcher Dr. Najib Ayas, Dr. Julie Carrier of the University of Montreal and investigator Dr. Allan Pack of the University of Pennsylvania, the study aims to identify a more precision approach to OSA treatment.
“We want to get to a point where we can analyze patients’ genetic and biochemical signatures to assess the severity of their sleep apnea and determine how aggressive the treatment course should be,” says Ayas.
“This would be a substantial paradigm shift from the current essentially one-size-fits-all approach, and would allow us to more precisely treat patients according to their OSA-related physiological vulnerabilities.”
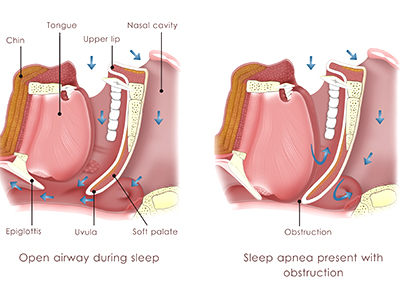
Over 900,000 Canadians have moderate to severe OSA, and many will be referred to a sleep clinic for assessment and treatment recommendations. However, current treatment approaches often target acute symptoms rather than the underlying causes and potential long-term consequences of OSA.
Notably, current OSA risk stratification diagnostics do not include biochemical indicators, such as inflammation and oxidative stress, that have been causally linked with OSA and OSA-related conditions. This, Ayas explains, is an information gap that his current research hopes to fill.
“We plan to use the patient data we are gathering to identify genetic and epigenetic signatures that might be associated with obstructive sleep apnea and the development of other conditions, for example, future OSA-associated cardiovascular or kidney disease.”
Tailoring sleep apnea treatments down to the level of a patient’s genetic code
Ayas’s study will draw from data collected through the CIHR-funded Canadian Sleep and Circadian Network (CSCN) Adult Obstructive Sleep Apnea Biobank. CSCN contains biological samples—blood samples, urine, etc.—along with demographic, cognitive, physiologic and medical history data from sleep clinic patients across Canada.
Ayas and his team will perform whole genome sequencing and DNA methylation on 1,500 patient samples. Both techniques are useful means to identify potential genetic precursors to OSA, along with other conditions.
Stanford University will contribute another approximate 1,500 analyses from their OSA patient biobank and genetic sequencing undertakings.

On top of this, Ayas and his team will cross-reference their samples with information found in provincial health databases across Canada, such as the Population Data BC (PopData) database, which house detailed patient data from publicly funded health services records.
“Once we connect our data to these data sets, we will be able to see how many patients with a certain genetic or epigenetic signature went on to have, for example, a heart attack or stroke, or were admitted to a care facility for dementia,” says Ayas.
“We will then be able to look at which parts of the genetic or epigenetic code predicted whether these patients would develop these other conditions.”
If a patient is found to be at greater risk of developing kidney failure or heart disease in the coming years, treatment programs could be specifically designed to target and prevent those ailments.
Ayas and his team will also use advanced signal processing techniques using polysomnography technology, which can connect unique relationships between cardiopulmonary (heart and lung) interactions, to see if these are predictive of future complications.
“If this research proves successful, we could one day provide patients with a more nuanced representation of their disease severity, and develop personalized care plans to try to prevent negative outcomes.”


